– Sidrat Tasawoor Kanth
A recent study from the Indian Institute of Science (IISc), published in the Journal of Proteome Research, has identified multiple mutations and unique proteins in isolates of SARS-CoV-2, the virus that causes COVID-19. It has also shown that the host produces several proteins of their own as their body launches an immunological defense in response to the viral attack.
COVID-19 has claimed over 2.5 million lives in just over a year. Humanity continues to face new challenges with novel strains – or genetic variants – of the virus being reported from around the world. To better understand how the virus is mutating and its protein biology (proteins are made using genetic information), an IISc team led by Utpal Tatu, Professor in the Department of Biochemistry, has carried out a comprehensive “proteo-genomic” investigation – a series of analyses of SARS-CoV-2 isolates. The isolates or viral samples were recovered from nasal secretions of consenting COVID-19 positive individuals in Bengaluru.
The genomic analysis was done using what molecular biologists like Tatu call next generation sequencing (NGS), a technology that allows for rapid sequencing of the entire genome. He says that sequencing the genomes of viral strains from around the world is important because it helps keep track of mutations that are arising constantly. His team’s analysis suggests that the virus is now mutating faster than before – the three Bengaluru isolates had 27 mutations in their genomes with over 11 mutations per sample, more than both the national average (8.4) and global average (7.3).
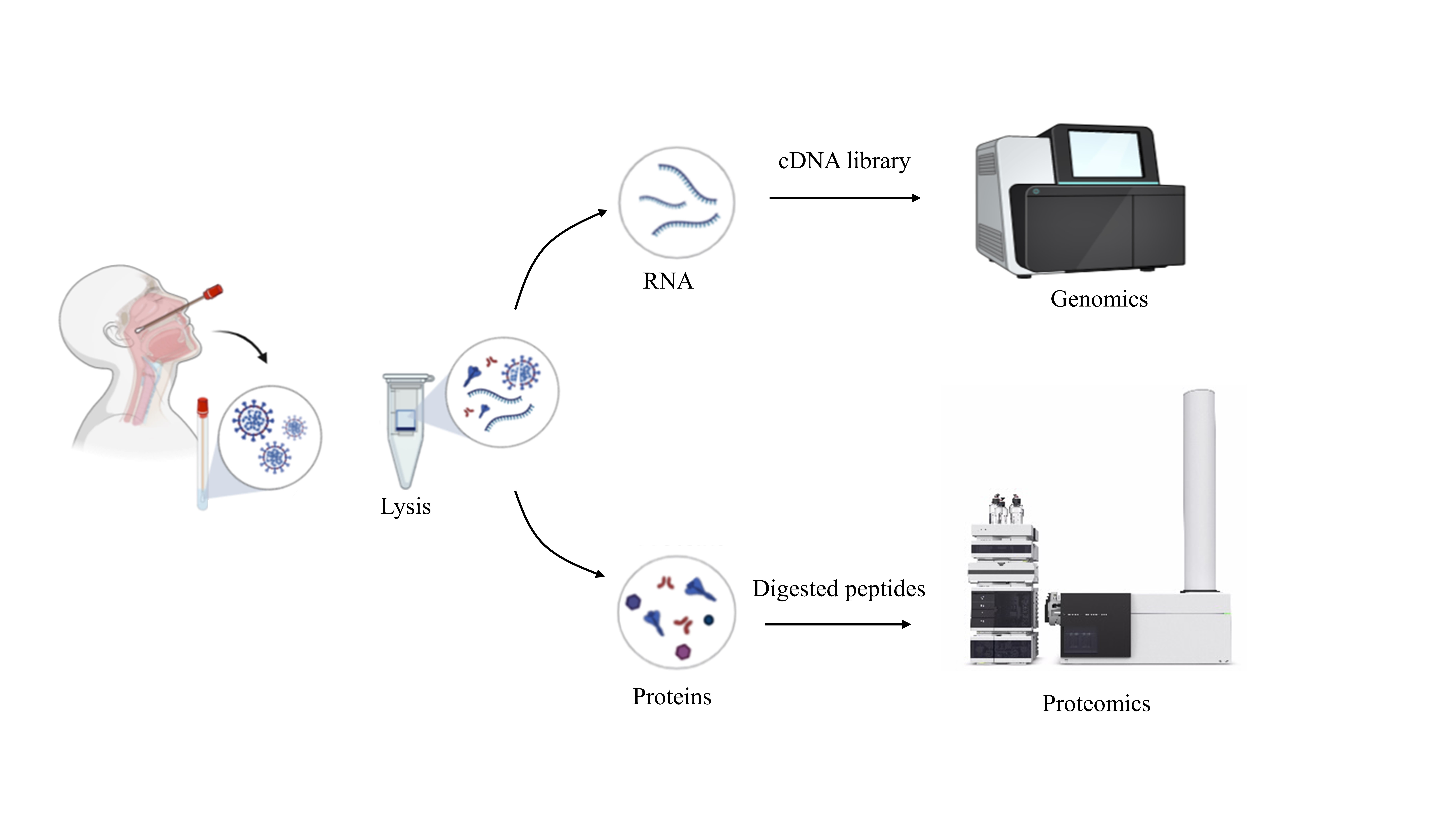
Steps involved in obtaining viral samples and performing proteo-genomic analysis (Credit: Sheetal Tushir)
To understand the spread and evolutionary history of the virus, the team constructed a global phylogenetic tree, or a tree of relatedness, of viral isolates using the sequence data. The phylogenetic analysis found that the Bengaluru isolates are most closely related to the one from Bangladesh. It also showed that the isolates in India have multiple origins rather than having evolved from a single ancestral variant.
The SARS-CoV-2 genome codes for more than 25 proteins, but only a handful of these proteins have been identified so far. “Studying viral proteins provides functional information which is currently not well represented,” says Tatu. In the proteomic analysis, his team detected 13 different proteins – most of them previously unidentified – from clinical samples. One such protein called Orf9b, which suppresses the host’s immune response, had been predicted, but the IISc team provided the first evidence of its expression.
“Just knowing how the virus functions will not be enough. We need to put it in the context of the host,” Tatu says. Therefore, in the third analysis, his team explored how our bodies respond to the virus by examining host proteins. They discovered as many as 441 proteins unique to COVID-19 positive patients, many of which are speculated to play a key role in the body’s immune response.
The proteomic analyses were carried out using a technique called high resolution mass spectrometry. The team is upbeat about the potential that this method has for large-scale testing. Proteins can be reliable markers of infections like COVID-19 because they are more abundant and stable as compared to RNA molecules on which the prevalent RT-PCR tests rely. Sheetal Tushir, a PhD student and the paper’s first author says, “The best thing we can [hope to] see in this century is the use of mass spectrometry as a basic technique for diagnostics.”
Tushir S, Kamanna S, Nath SS, Bhat A, Rose S, Aithal AR, Tatu U. Proteo-Genomic Analysis of SARS-CoV-2: A Clinical Landscape of Single-Nucleotide Polymorphisms, COVID-19 Proteome, and Host Responses. J Proteome Res. 2021
https://pubs.acs.org/doi/abs/10.1021/acs.jproteome.0c00808
CONTACT:
Utpal Tatu
Professor
Department of Biochemistry
Indian Institute of Science (IISc)
tatu@iisc.ac.in
+91-80-2293 2823
NOTE TO JOURNALISTS:
a) If any of the text in this release is reproduced verbatim, please credit the IISc press release.
b) For any queries about IISc press releases, please write to news@iisc.ac.in or pro@iisc.ac.in



